Supporting Information
Supporting Information
Eric R Bittner
Department of Physics, University of Houston
Algorithmic preliminaries
Algorithmic preliminaries
(*============================================================DimerLiouvillian(full16x16)--trial-runscaffold============================================================*)ClearAll[c,LfunRelax,Lrelax,kplus,kminus,Lplus,Lminus,LD,LH];(*----------basic2-leveloperators----------*)id2=IdentityMatrix[2];sp=SparseArray[{{2,1}->1},{2,2}];(*sigma_+*)sm=SparseArray[{{1,2}->1},{2,2}];(*sigma_-*)sz=SparseArray[{{1,1}->1,{2,2}->-1},{2,2}];(*two-spinembedding*)op1[A_]:=KroneckerProduct[A,id2];op2[A_]:=KroneckerProduct[id2,A];sm1=op1[sm];sm2=op2[sm];sp1=op1[sp];sp2=op2[sp];sz1=op1[sz];sz2=op2[sz];id4=IdentityMatrix[4];(*----------vectorizationconvention----------vec(ArhoB)=(Transpose[B]⊗A)vec(rho)------------------------------------------------*)LH[H_]:=-I(KroneckerProduct[id4,H]-KroneckerProduct[Transpose[H],id4]);LD[L_]:=Module[{LdL=ConjugateTranspose[L].L},KroneckerProduct[Conjugate[L],L]-1/2KroneckerProduct[id4,LdL]-1/2KroneckerProduct[Transpose[LdL],id4]];
MODEL A : correlated relaxation (collective decay)
MODEL A : correlated relaxation (collective decay)
Take =(1)±(2)) and Hamiltonian : detuning in single - excitation manifold (| 01 > vs | 10 >) such that =(|01〉〈01|-|10〉〈10|) is diagonal in the in the computational basis
L
±
γ(1±c
(-
σ
-
σ
2
H
Δ
Δ
2
proj01=SparseArray[{{2,2}->1},{4,4}];proj10=SparseArray[{{3,3}->1},{4,4}];HDelta[Delta_?NumericQ]:=(Delta/2)(proj01-proj10);Lplus=(sm1+sm2)/Sqrt[2];Lminus=(sm1-sm2)/Sqrt[2];kplus[gamma_?NumericQ,c_?NumericQ]:=gamma(1+c);kminus[gamma_?NumericQ,c_?NumericQ]:=gamma(1-c);Lrelax[Delta_?NumericQ,gamma_?NumericQ,c_?NumericQ]:=LH[HDelta[Delta]]+kplus[gamma,c]LD[Lplus]+kminus[gamma,c]LD[Lminus];
MODEL B:correlated dephasing (collective sigma_z noise)
MODEL B:correlated dephasing (collective sigma_z noise)
Take =(1)±(2)) . Choose an SA-basis tunneling Hamiltonian equivalent in {|01>,|10>} and coupling/tunneling via the hamiltonian =J(|01〉〈10|+|10〉〈01|) in the computational basis.
L
±
γ(1±c)
(σ
z
σ
z
2
H
J
hop0110=SparseArray[{{2,3}->1,{3,2}->1},{4,4}];(*SAFENAME*)HJ[J_?NumericQ]:=Jhop0110;Zplus=(sz1+sz2)/Sqrt[2];Zminus=(sz1-sz2)/Sqrt[2];Ldeph[J_?NumericQ,gamma_?NumericQ,c_?NumericQ]:=LH[HJ[J]]+kplus[gamma,c]LD[Zplus]+kminus[gamma,c]LD[Zminus];
EP Diagnostics
EP Diagnostics
(*============================================================EPdiagnostics(simple+robust)-"near-degeneracy"ineigenvalues(withineps)-rankdropofeigenvectormatrix(Jordandefectproxy)============================================================*)clusterEigenvalues[evs_List,eps_?NumericQ]:=Module[{clusters={},used,n=Length[evs]},used=ConstantArray[False,n];Do[If[used[[i]],Continue[]];Module[{cl={i},changed=True},used[[i]]=True;While[changed,changed=False;Do[If[used[[j]],Continue[]];If[Min[Abs[evs[[j]]-evs[[#]]]]&/@cl//Min<eps,AppendTo[cl,j];used[[j]]=True;changed=True;],{j,1,n}];];AppendTo[clusters,cl];],{i,1,n}];clusters];epFlagByRank[Lmat_?MatrixQ,epsEV_:10^-6,tolRank_:10^-8]:=Module[{evs,vecs,clusters,ranks,maxCl,rankV},{evs,vecs}=Eigensystem[Lmat];clusters=clusterEigenvalues[evs,epsEV];(*focuson“largest”clusterasEPcandidate*)maxCl=First@MaximalBy[clusters,Length];rankV=MatrixRank[Transpose[vecs[[maxCl]]],Tolerance->tolRank];(*EPcandidateifclustersize>rank(defect)*)If[Length[maxCl]>rankV,1,0]];
In[]:=
(*---EPstrengthviaeigenvectorconditioning---*)ClearAll[epStrength,epFlag];epStrength[L_?MatrixQ,tolSV_:10^-12]:=Module[{vals,vecs,V,smin},{vals,vecs}=Eigensystem[N[L,50]];V=Transpose[vecs];(*columns=eigenvectors*)smin=Min[SingularValueList[V]];1/Max[smin,tolSV]];(*turnstrengthintoabinaryflagifyouwant*)epFlag[L_?MatrixQ,thresh_:10^6]:=If[epStrength[L]>thresh,1,0];
Test cases
Test cases
This generates the graphics in the manuscript. Ideally, the end user should be able to plug-and-play with their own Liouvillian super-op “flattened” into a x array.
2
d
2
d
In[]:=
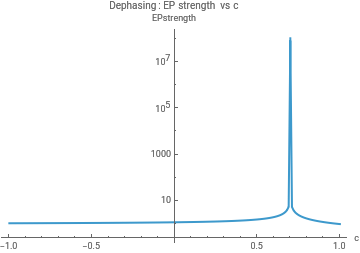
cGrid=Range[-1,1,0.01];(*chooseONE:Lfun=LdephorLrelax*)params=<|"gamma"->1,"Delta"->0.5,"J"->0.3|>;LfunDeph[c_]:=Ldeph[params["J"],params["gamma"],c];LfunRelax[c_]:=Lrelax[params["Delta"],params["gamma"],c];dataDephStrength=Table[{c,epStrength[LfunDeph[c]]},{c,cGrid}];dataRelaxStrength=Table[{c,epStrength[LfunRelax[c]]},{c,cGrid}];ListLogPlot[dataDephStrength,Joined->True,PlotRange->All,AxesLabel->{"c","EPstrength"},PlotLabel->"Dephasing: EP strength vs c"]ListLogPlot[dataRelaxStrength,Joined->True,PlotRange->All,AxesLabel->{"c","EPstrength"},PlotLabel->"Relaxation: EP strength vs c"]params
Out[]=