Modeling Enzyme Efficiency with an Inhibitor-Adjusted Michaelis–Menten Equation
Modeling Enzyme Efficiency with an Inhibitor-Adjusted Michaelis–Menten Equation
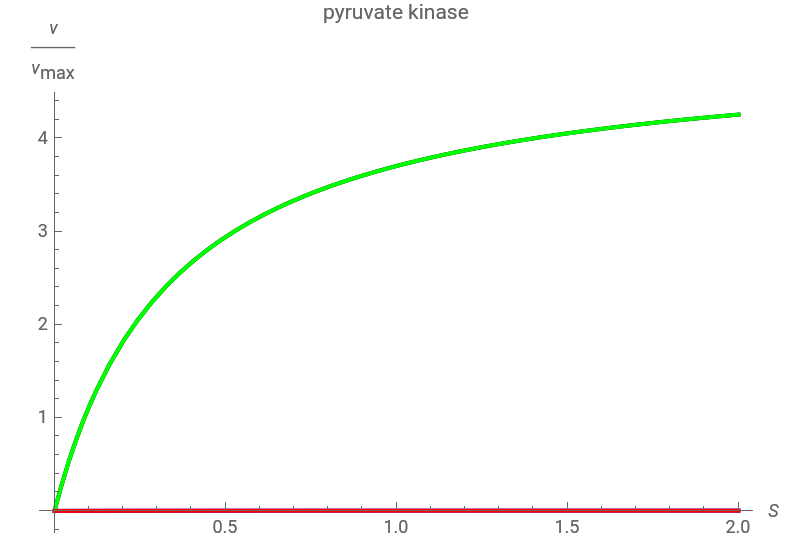
This Demonstration considers enzyme efficiency in environments of changing substrate, competitive inhibitors and allosteric inhibitors. Using the Michaelis–Menten equation, the plot shows the reaction rate based on substrate concentration and the relative concentrations of inhibitors. Variables like the maximum reaction rate and the Michaelis constant are intensive enzyme properties used to describe the reaction rates of pyruvate kinase and Rubisco.
Details
Details
Enzymes dictate the world around us, and their importance cannot be understated. For example, Ribulose-1,5-bisphosphate carboxylase/oxygenase (known as Rubisco) is largely responsible for the integrity of the entire food chain and almost all of the ecological diversity in the world today. Without one simple enzyme, life would be reduced to prokaryotes and fungi, only some of which are large enough for the human eye to see. Understanding and optimizing the efficiency of an enzyme like Rubisco would have incredible ramifications, including producing food more efficiently. Of course, Rubisco is only one enzyme of many, but it quickly becomes difficult to conceptualize the exceedingly high level of importance enzymes have and the outcome of optimizing their functions.
Click one of the two enzymes provided. Then drag the sliders to vary the concentrations of the competitive and noncompetitive inhibitor substrates to observe how the reaction rate changes across the entire range. You can also adjust the range of substrate concentration by using the "maximum" slider.
[S]
The Michaelis–Menten equation is used to visualize the data. This equation defines a reaction rate for a given enzyme with respect to substrate concentration and the maximum reaction rate . The Michaelis constant represents the substrate concentration at which the reaction rate is one-half of :
[S]
V
max
K
m
V
max
V=[S]+[S]
V
max
K
m
Since and depend on the enzyme selected, only two reactions are considered: the final step of glycolysis catalyzed by pyruvate kinase and the respiration catalyzed by Rubisco. Obviously, their substrates are not present in pure form in any natural settings, so competitive and noncompetitive inhibitor substrates affect how the reactions proceed. Their influences are defined by the following equations, where is the noncompetitive inhibitor constant, is the competitive inhibitor constant, is the noncompetitive inhibitor substrate concentration and is the competitive inhibitor substrate concentration:
V
max
K
m
K
n
K
c
I
1
I
2
V
noncompetitive
V
max
1+(+[S])
I
1
K
n
K
m
V
competitive
V
max
1+(+[S])
I
2
K
c
K
m
V
mixed
V
max
1+1++[S]
I
1
K
n
K
m
I
2
K
c
The kinetic constants are empirically calculated based on their most representative substrates. The enzyme concentration is an arbitrary number based on the referenced literature.
Rubisco:
V
max
K
m
K
c
-1
sec
Pyruvate kinase:
V
max
K
m
K
n
=6.2×M
-6
10
K
c
=980×M
-6
10
References
References
[1] H. Farazdaghi, "The Single-Process Biochemical Reaction of Rubisco: A Unified Theory and Model with the Effects of Irradiance, and Rate-Limiting Step on the Kinetics of C3 and C4 Photosynthesis from Gas Exchange," Biosystems, 103(2), 2011 pp. 265–284. doi:10.1016/j.biosystems.2010.11.004.
CO
2
[2] A. U. Igamberdiev and M. R. Roussel, "Feedforward Non-Michaelis–Menten Mechanism for Uptake by Rubisco: Contribution of Carbonic Anhydrases and Photorespiration to Optimization of Photosynthetic Carbon Assimilation," Biosystems, 107(3), 2012 pp. 158–166. doi:10.1016/j.biosystems.2011.11.008.
CO
2
[3] Uri M, "Kcat of RuBisCO." (Dec 17, 2024) bionumbers.hms.harvard.edu/bionumber.aspx?s=n&v=0&id=107429.
[4] B. Baranowska and T. Baranowski, "Kinetic Properties of Human Muscle Pyruvate Kinase," Molecular and Cellular Biochemistry, 45(2), 1982 pp. 117–125. doi:10.1007/BF00223506.
[5] BRENDA, "EC 2.7.1.40: pyruvate kinase." (Dec 17, 2024) www.brenda-enzymes.org/all_enzymes.php?ecno=2.7.1.40&table=KM_Value.
[6] M. Yuan et al., "An Allostatic Mechanism for M2 Pyruvate Kinase as an Amino-Acid Sensor," Biochemical Journal, 475(10), 2018 pp. 1821–1837. doi:10.1042/BCJ20180171.
[7] D. S. Y. Teo, S. F. Chew and Y. K. Ip, "L-Cysteine Is a Competitive Inhibitor of Pyruvate Kinase from the Intertidal Sipunculan, Phascolosoma arcuatum," Zoological Science, 17(6), 2000 pp. 717–723. doi:10.2108/zsj.17.717.
Permanent Citation
Permanent Citation
Elizabeth Chang, Siyi Du, Will Dulaney, Yunhan Fang, Harshini Rubarajan and Shreya Pillai
"Modeling Enzyme Efficiency with an Inhibitor-Adjusted Michaelis–Menten Equation"
http://demonstrations.wolfram.com/ModelingEnzymeEfficiencyWithAnInhibitorAdjustedMichaelisMent/
Wolfram Demonstrations Project
Published: December 17, 2024